A Case Study on 16s rRNA Sequencing of Fuel
A snapshot from global scenario
Microbial contamination of hydrocarbon fuel is a severe problem that can lead to costly and dangerous operational problems in fuel storage and engine systems [1]. Bacteria and fungi can cause deterioration of fuels through a number of mechanisms, including an accumulation of biomass, degradation of protective additives, and production of corrosive metabolic byproducts [2].
Identification of hydrocarbon fuel contaminating organisms depends heavily on culture based testing and traditional phenotypic identification [3,4]. Culture based methods may often lead to underestimation of contaminant diversity and misidentification of the constituent species [1]. The diagnosis may often be hindered by uncultivable or fastidious contaminants and usually requires about 7 days for the identification of the contaminants that are cultivable. In addition, efficient classification of contaminating taxa, particularly those which serve as early indicators of serious biofouling issues such as microbially influenced corrosion caused by sulfate-reducing bacteria (SRBs) [5,6] or fouling of engine components caused by biofilm-producing bacteria may prompt early intervention [1].
 Direct application of 16s rRNA gene sequencing in hydrocarbon fuel contamination testing has massively changed the field of microbial ecology, and has a great potential to provide new insights into the microbiology of fuel contamination [1]. Massively parallel next generation sequencing of 16s rRNA gene has enabled to assess the diversity the whole spectrum of microbes, including the uncultivable taxa at a deeper level.
Direct application of 16s rRNA gene sequencing in hydrocarbon fuel contamination testing has massively changed the field of microbial ecology, and has a great potential to provide new insights into the microbiology of fuel contamination [1]. Massively parallel next generation sequencing of 16s rRNA gene has enabled to assess the diversity the whole spectrum of microbes, including the uncultivable taxa at a deeper level.
16s rRNA sequencing of bacteria is the current most successful method for bacterial identification. It is a very rapid technique with only 48 hours of operational time from start to end, and the bacteria can be characterized without the need for clone library generation. The technique can discriminate between closely related isolates, and can detect their relative abundances with high accuracy. These advantages allow early detection of microbial influenced biofouling issues, which permits the issues to be addressed faster, when they are still underway.
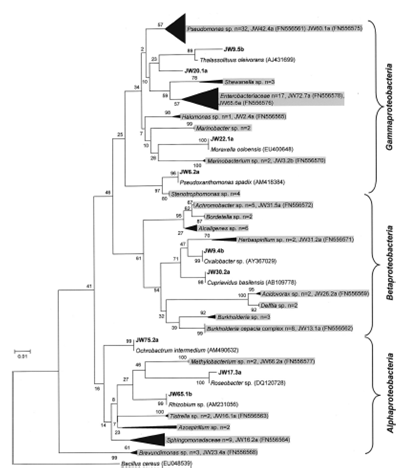
Fig- Gram negative bacteria isolated from contaminated fuel samples [1]
Need of implementing microbial quality assurance of fuel in Sri Lanka
Although disregarded, there prevails a timely need of establishing a system for microbial quality assessment of hydrocarbon fuel used within the country. According to a World Bank report published in 2010, the road sector gasoline fuel consumption (kt of oil equivalent) in Sri Lanka was 452 in 2009 [7] and the crude oil consumption reached up to 97,000 barrels per day in 2011[8]. Having a vast array of logistics and transportation agencies being operated in the country, Sri Lanka requires a rapid, accurate and scalable crude oil quality control system in microbiological aspects. Biofouling of fuel may lead to serious, costly and dangerous issues related to operation failures of engines, low efficiency due to incomplete combustion and environmental pollution.
To address the gap, Credence Genomics has introduced a rapid, accurate and highly scalable microbial testing technique of 16s rRNA Sequencing, which is a leading edge technology adopted as favorite in laboratories worldwide, facilitating a newer era of laboratory testing.
To summarize, 16s rRNA Sequencing has following advantages compared to other techniques-
| Conventional techniques | 16s rRNA Sequencing |
|---|---|
| Longer turnaround times. A single test requires a minimum of 7 days | Lesser turnaround times. Extremely rapid with a run time of only 12 hours. |
| Test accuracy is dependent on operator expertise. The accuracy of testing Is,̴80%[9] | Accuracy is operator-independent. Highly accurate with an accuracy of 99.9% |
| Fastidious, unculturable organisms and unusual biochemical reactions impede the identification process. | Test method is culture independent; Growth requirements, biochemical reactions and culture failures do not affect the identification process. |
| Only a limited number of organisms can be identified. Important organisms in a test sample can be missed out. | Identifies the complete range of organisms present in a test sample. |
| The percentage of strains correctly identified to the species level is sometimes less than satisfactory. | Permits identification of complete spectrum of organisms down to species level, their intragenic variations and relative abundances |
| Limited throughput, Sensitivity may vary. | Dramatic increase in throughput; High sensitivity |
References-
- Judith White, Jack Gilbert, Graham Hill, Edward Hill, Susan M. Huse,Culture-Independent Analysis of Bacterial Fuel Contamination Provides Insight into the Level of Concordance with the Standard Industry Practice of Aerobic Cultivation
- Allsopp, Dennis. Introduction to biodeterioration. Cambridge University Press, 2004.
- Institute of Petroleum. Determination of the viable aerobic microbial content of fuels and fuel components, p. 385.1-385.7. In Standard methods for the analysis and testing of petroleum products and British standard 2000 Parts. Energy Institute, London, United Kingdom.
- Institute of Petroleum. Guidelines for the investigation of the microbial content of petroleum fuels and for the implementation of avoidance and remedial strategy, 2nd ed. The Energy Institute, London, United Kingdom.
- Brown, L. M., et al. “Community dynamics and phylogenetics of bacteria fouling Jet A and JP-8 aviation fuel.” International Biodeterioration & Biodegradation3 (2010): 253-261.
- Campbell, Maureen, Eshwar Mahenthiralingam, and David P. Speert. “Evaluation of random amplified polymorphic DNA typing of Pseudomonas aeruginosa.” Journal of clinical microbiology12 (2000): 4614-4615.
- com. 2013. SRI LANKA | ECONOMIC INDICATORS | DATA LIST BY COUNTRY. [online] Available at: http://www.tradingeconomics.com/sri-lanka/ [Accessed: 23 Nov 2013].
- Sri Lanka Crude Oil Consumption by Year
- Janda, J. Michael, and Sharon L. Abbott. “Bacterial identification for publication: when is enough enough?.” Journal of clinical microbiology6 (2002): 1887-1891.


