Performance Evaluation of Sewage Treatment Plants in Sri Lanka
Situation
Sri Lanka confronts a variety of water and wastewater issues and water related health hazards [1]. These problems arise primarily due to the increasing level of qualitative and quantitative depletion of water resources owing to over- utilization and continuous discharge of wastewater. More than 90% of sewerage in the developing world is discharged directly into rivers, lakes, and coastal waters without any treatment. Large cities such as Colombo, Galle, Jaffna and Kandy have severe problems disposal of sewerage, industrial effluents and industrial and of domestic solid waste, as they generate large quantities but have no facilities for their treatment and proper disposal [2]. UNEP (2001) estimated that each day in the Colombo Metropolitan Region, 428 MT of sewerage is released into the ground through septic tanks and pit latrines and 138 MT of sewerage is released into waterways [3]. In central province Kandy suffers from serious problems of wastewater disposal, which majorly contributes to the pollution of Mahaweli river. No proper system is been established for wastewater disposal in Kandy, and about 80% of used water is released as wastewater [2]. In addition, Pussalla Oya catchment was the main source area for the “Hepatitis A” outbreak recorded in Gampola district in May 2007 [4]
To reduce the degradation of water quality and ensure a healthy environment, Sewage Treatment Plants have been constructed in many areas, especially in rapidly urbanizing cities around the country. However water treatment efficacy has been a major concern [2] Different authorities concerning public health have set up guidelines on maximum allowable microbial, physical and chemical concentrations in treated sewage water. There are currently several approaches in case of establishing microbiological quality of treated water, based on conventional testing methods. Routine culture based testing is performed to identify bacterial pathogens; Vibro cholera, Salmonella typhi, other Salmonella spp, Shigella spp., Ca mphylobacter, Proteus spp., Coiform spp.,E.coi 0157, Clostridium spp., Pseudomonas spp., Tuberclebacilli, Leptospira and Yersinia enterocolitica that are the main causative agents of human gastro intestinal diseases[5]. Culture based techniques always depend upon index organisms and indicator parameter testing which usually takes about a week. Some fastidious bacteria would show delayed or unsatisfactory growth patterns that usually obstruct the identification process, and anaerobic bacterial groups can be specifically hard to identify. In addition to these, culturing techniques do not facilitate intragenic variation detection in a sample containing a mixed population. Thus, an efficient procedure for treated sewage testing is a timely need as poorly treated wastewater, when used for agricultural, recreational, and other purposes can act as the key for many contagious diseases.
 Solution
Solution
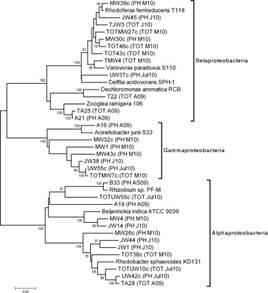
Fig-Phylogenetic analysis of 16S rRNA gene sequences of Proteobacteria from the oxidation tank of a wastewater treatment plant [6]
16s rRNA gene sequencing is performed worldwide as a rapid and reliable technique for the determination of the efficiency of waste water treatment. This is a very cost-effective method, large scale method for assessing microbial composition, their relative abundances and intragenic variations. In addition, it helps to prevail over major drawbacks of culture based techniques in terms of culture failures, unculturable organisms, and time as 16s rRNA sequencing is a culture independent method. The total profile of microorganisms in the sample can be identified within just 24 hours of runtime, thus enabling a remarkable increase the throughput. This can be used as the best solution to establish high standards of water treatment processes to ensure better safety and microbiological quality of treated water.
To summarize, 16s rRNA Sequencing has following advantages compared to other techniques-
| Conventional techniques | 16s rRNA Sequencing |
|---|---|
| Longer turnaround times. A single test requires a minimum of 7 days | Lesser turnaround times. Extremely rapid with a run time of only 12 hours. |
| Test accuracy is dependent on operator expertise. The accuracy of testing Is,̴80%[9] | Accuracy is operator-independent. Highly accurate with an accuracy of 99.9% |
| Fastidious, unculturable organisms and unusual biochemical reactions impede the identification process. | Test method is culture independent; Growth requirements, biochemical reactions and culture failures do not affect the identification process. |
| Only a limited number of organisms can be identified. Important organisms in a test sample can be missed out. | Identifies the complete range of organisms present in a test sample. |
| The percentage of strains correctly identified to the species level is sometimes less than satisfactory. | Permits identification of complete spectrum of organisms down to species level, their intragenic variations and relative abundances |
| Limited throughput, Sensitivity may vary. | Dramatic increase in throughput; High sensitivity |
References-
- Well Rainwater, S. and Dug, P. 2006. The Global Water Supply and Sanitation Assessment 2000 Report (WHO and UNICEF 2000), the most recent compilation of global statistics on water supply, changed the way that such data are compiled, from the previous unreliable estimates by provider agencies to consumers’ responses in population-based. A custom publication of the Disease Control Priorities Project, p. 114-126.
- Sudasinghe, M. I., L. W. Galagedara, and E. R. N. Gunawardena. “Performance Evaluation of Selected Sewerage Treatment Plants in Sri Lanka.”Tropical Agricultural Research 22.2 (2011): 154-164.
- (2001). Annual report, Regional Resource Centre for Asia and the Pacific 2001, Sri Lanka: State of the environment 2001. Pathumthani, Thailand: UNEP.
- Abeysinghe, M.R.N. (2007). Weekly Epidemiological Report, Ministry of Healthcare and Nutrition, Government of Sri Lanka. 15-21.
- Hoogenboezem(2007);Influences of sewage treatment plant effluents on the occurrence of emerging waterborne pathogens in surface water
- Del Casale, Antonio, et al. “Extent and variation of phage-borne bacterial 16S rRNA gene sequences in wastewater environments”.
Applied and environmental microbiology 77.15 (2011): 5529-5532. - Janda, J. Michael, and Sharon L. Abbott. “Bacterial identification for publication: when is enough enough?.” Journal of clinical microbiology 40.6 (2002): 1887-1891.


