Situation
The microbial safety of cosmetic and personal care products is an important consideration.
About 30% of cosmetics currently sold in supermarkets, pharmacies and cosmetic shops do not have a proper permit from the Cosmetics, Devices and Drugs Regulatory Authority [1]. Recently, newspapers reported a death of a woman caused due to an anaphylaxis reaction triggered by using a low quality cosmetic product [2]. Therefore, it is essential to maintain the established standards of quality of cosmetic and personal care products, as poor quality may lead to adverse health effects or even death.
Current microbiological practices used in microbial quality testing of cosmetics and personal care products involve culture plating for enumeration and identification purposes. Mainly, presence or absence of bacteria such as Staphylococcus aureus, Pseudomonas aerogenosa and E.coli is tested using identification methods [3].Although these tests permit identification of contaminants, they require a minimum turnaround time of 7 days, and the identification can always be impeded by non-culturable or fastidious strains. Such culture based techniques are always biased towards fast growing populations. In addition, the number of tests carried out to determine possible fungal contaminations are inadequate. Therefore, the product’s hygiene with respect to the possibility of fungal contamination can be questionable. The long time required for microbial identification using culture based methods may intervene with the efficiency of the manufacturing process.
 Solution
Solution
Next-Generation-Sequencing based identification of bacteria and fungi is adopted in laboratories worldwide as a very successful method for detailed identification of microbial contaminations.
It involves rapid sequencing of 16s rRNA gene of bacteria and ITS region of fungi, and only requires 48 hours of runtime, accelerating the identification process as well as the manufacturing efficiency. Since 16s rRNA gene/ITS region sequencing is a culture independent method, it allows identification of whole the spectrum of microorganisms that could be present in a test sample. The technique is capable of testing up to 200 samples in a single run, generating highly accurate results. Not only can this method identify organisms up to species level, but it can also discriminate between closely related species as well as their relative abundances permitting highly sensitive and accurate identification of both bacteria and fungi. Therefore, the technique can be an advantage in many ways, from better, faster and accurate identification of contaminants to increased production efficiency.
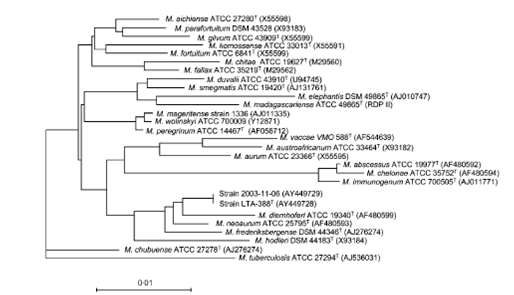
In addition to industrial applications, 16s rRNA gene/ ITS gene sequencing can be used to assess cosmetic clinics for environmental contaminants and disinfection efficiencies or even to test isolates from cosmetic infection sites. Using 16s rRNA sequencing of bacteria, a recent study identified a novel, rapidly growing Mycobacterium spp, Mycobacterium cosmeticum, from a patient with a cosmetic infection [4].
Therefore, 16s rRNA gene/ ITS gene sequencing can be applied as a comprehensive approach to conduct detailed studies on harmful microbial contaminants of cosmetics and personal care products. Having such a rapid, accurate, sensitive technique within reach, manufacturers can assess and assure the quality of their products, as well as their cGMP (current Good Manufacturing Practices) in order to maintain higher production efficiencies.
To summarize, 16s rRNA Sequencing has following advantages compared to other techniques-
| Conventional techniques | 16s rRNA Sequencing |
| Longer turnaround times. A single test requires a minimum of 7 days | Lesser turnaround times. Extremely rapid with a run time of only 12 hours. |
| Test accuracy is dependent on operator expertise. The accuracy of testing Is low( ̴80%)[5] | Accuracy is operator-independent. Highly accurate with an accuracy of 99.9% |
| Fastidious, unculturable organisms and unusual biochemical reactions impede the identification process. | Test method is culture independent; Growth requirements, biochemical reactions and culture failures do not affect the identification process. |
| Only a limited number of organisms can be identified. Important organisms in a test sample can be missed out. | Identifies the complete range of organisms present in a test sample. |
| The percentage of strains correctly identified to the species level is sometimes less thansatisfactory. | Permits identification of complete spectrum of organisms down to species level, their intragenic variations and relative abundances |
| Limited throughput, Sensitivity may vary. | Dramatic increase in throughput; High sensitivity |
References-
- Manjula Fernando. 2011. Beauty – at what price?. [ONLINE] Available at:http://www.sundayobserver.lk/2011/10/23/fea09.asp. [Accessed 18 September 13].
- Nadia Fazlulhaq. 2011. Beauty treatment ends in tragedy. [ONLINE] Available at:http://www.sundaytimes.lk/111016/News/nws_21.html. [Accessed 18 September 13].
- Sri Lanka Standards Institution . 2011. Laboratory Services . [ONLINE] Available at:http://www.slsi.lk/web/index.php?option=com_content&view=article&id=80&Itemid=98&lang=en. [Accessed 19 July 13]
- Cooksey, Robert C., et al. “Mycobacterium cosmeticum sp. nov., a novel rapidly growing species isolated from a cosmetic infection and from a nail salon.” International journal of systematic and evolutionary microbiology6 (2004): 2385-239
- Janda, J. Michael, and Sharon L. Abbott. “Bacterial identification for publication: when is enough enough?.” Journal of clinical microbiology6 (2002): 1887-1891.


