A case study on ecological microbial diversity and its importance
Situation
Microorganisms constitute most of the biodiversity on earth, and they are predominant in every ecosystem found in the environment. Several of the processes which microorganisms perform are of critical importance for the cycling of nutrients, the degradation of various compounds, and the global climate [1]. Knowledge of microorganisms in environment; mainly in rainforests, freshwater and marine ecosystems in Sri Lankan context help mankind to develop services and find strategies to utilize natural resources in a sustainable manner as well as to conserve the ecological diversity for future generations.
In their ecological habitats, both bacteria and fungi act as biodegrading agents of decaying materials which helps in recycling of nutrients and maintenance of nutrient cycles. With technological revolution the global environmental microbiology has undergone over the past decade [1], this natural phenomenon is used for the benefit of mankind in finding sustainable ways to clean up contaminated environments [2]. Therefore, a deeper knowledge on ecological microorganisms will open doors to an entirely new world that will help to reduce environmental pollution in an efficiently economical manner.
Bioremediation or the introduction of group of natural microbial strains or genetically engineered variants to reinforce the treatment of contaminated environments, can be used effectively to treat wastewater [2] and remove treat effluents of different industries such as pharmaceuticals, stainless steel plants, preparation of alloys, chrome plating, leather tanning, production of refractories, dye industry, industrial water cooling, paper pulp production, petroleum refining, wood preservation [3] as well as to remove oil spills from oceans [4] and to address the deteriorating soil conditions [5]. It is a highly efficient, less expensive, highly specific natural process which results in harmless residues that carbon dioxide, water, and cell biomass. Bioremediation allows complete destruction of target population instead of transferring contaminants from one environmental medium to another [6].
Some companies in Sri Lanka have already established this landmark approach [5], but the process can be enhanced by widening the knowledge on ecological microorganisms that efficiently act as bioremediation agents and by provision of newer techniques that can overcome the limitations of current conventional and molecular biological techniques used in identification of microorganisms.
Current conventional methods used in microbiological identification include culturing of bacteria and fungi, but the inability to culture certain species of microbes intrude the identification process. Culture based methods are bias towards the fast growing species in a population, and requires a minimum turnaround time of 7 days. Although Density Gradient Gel Electrophoresis (DGGE) can be used for microbial identification, the phylogenetic relations established using the technique can be less reliable, in turn reducing the sensitivity of the identification. Fluorescent In Situ Hybridization (FISH) can also be used as a molecular based technique, but a prior knowledge of the ecosystem under study and the microorganisms most likely to be detected is necessary for the process. In addition, preparation of probes can be an expensive, cumbersome and unsatisfactory [7].
Solution
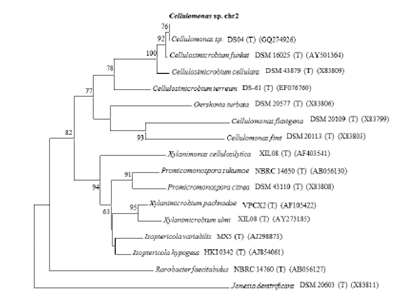 Fig- phylogenetic tree based on 16s rRNA gene sequence comparisons showing the position of isolate Cellulomonas spp [3].
Fig- phylogenetic tree based on 16s rRNA gene sequence comparisons showing the position of isolate Cellulomonas spp [3].
Next generation sequencing of 16s rRNA of bacteria and ITS region of fungi is one of the rapid, most accurate and comprehensive techniques used to eliminate the barriers of conventional techniques in discovering ecologically important strains of microorganisms as well as their functions in ecosystems, in order to utilize them in an efficient manner. This novel technique is culture independent, and requires only very minute quantities of test organism DNA (20 ng).
Having the fastest runtime ever known to a molecular based method, it only requires 48 hours to perform a run parallelized up to 200 test samples. Sequencing of 16s rRNA of bacteria and ITS region of fungi is a very sensitive identification approach, which identifies both bacteria and fungi up to species level while discriminating between the intragenic variations of closely related species. It can also provide measurements on relative abundances and species richness of test samples.
Therefore, sequencing of 16s rRNA of bacteria and ITS region of fungi provides new insights to the field, accelerating new discoveries that may contribute towards finding of sustainable strategies of diversity conservation, removal of pollutants, effluent treatment and many more.
To summarize, 16s rRNA Sequencing has following advantages compared to other techniques-
| Conventional techniques | 16s rRNA Sequencing |
|---|---|
| Longer turnaround times. A single test requires a minimum of 7 days | Lesser turnaround times. Extremely rapid with a run time of only 12 hours. |
| Test accuracy is dependent on operator expertise. The accuracy of testing Is,̴80%[9] | Accuracy is operator-independent. Highly accurate with an accuracy of 99.9% |
| Fastidious, unculturable organisms and unusual biochemical reactions impede the identification process. | Test method is culture independent; Growth requirements, biochemical reactions and culture failures do not affect the identification process. |
| Only a limited number of organisms can be identified. Important organisms in a test sample can be missed out. | Identifies the complete range of organisms present in a test sample. |
| The percentage of strains correctly identified to the species level is sometimes less than satisfactory. | Permits identification of complete spectrum of organisms down to species level, their intragenic variations and relative abundances |
| Limited throughput, Sensitivity may vary. | Dramatic increase in throughput; High sensitivity |
Reference-
- Sustanability formas.se. 2013. Issue #3, September 2013 – Sustainability. [online] Available at: http://sustainability.formas.se [Accessed: 23 Nov 2013].
- Vivehananthan, K., Y. W. Manukulasooriya, and W. M. J. Y. Ekanayake. “Detection of Potential Eco-friendly Microbes to Clean-Up the Polluted Water in Sri Lanka.” Journal of Environmental Professionals Sri Lanka 1.2 (2013): 76-82.
- Focardi, Silvia, Milva Pepi, and Silvano E. Focardi. “Microbial Reduction of Hexavalent Chromium as a Mechanism of Detoxification and Possible Bioremediation Applications.” (2013).
- 2013. [online] Available at: 4. http://resources.scienceworld.ca/demonstration/bioremediation-of-oil-spill [Accessed: 23 Nov 2013]
- org. 2013. Bioremediation at Dilmah tea gardens | Dilmah Conservation. [online] Available at: http://www.dilmahconservation.org/initiatives/bio-remediation-at-dilmah-tea-gardens/ [Accessed: 23 Nov 2013].
- Vidali, M. “Bioremediation. An overview.” Pure and Applied Chemistry 73.7 (2001): 1163-1172.
- Sanz, Jose L., and Thorsten Köchling. “Molecular biology techniques used in wastewater treatment: an overview.” Process Biochemistry 42.2 (2007): 119-133.
- Janda, J. Michael, and Sharon L. Abbott. “Bacterial identification for publication: when is enough enough?.” Journal of clinical microbiology 40.6 (2002): 1887-1891.


